In Lead Molecular Design we develop scientific software in colaboration with Molecular Discovery Ltd.. Our main products are listed below, including the MS file converter to translate the binary files from Waters, Bruker and Thermo Fisher Scientific LCMS systems into a SQLite file format that can be used by MassMetaSite and MassChemSite.
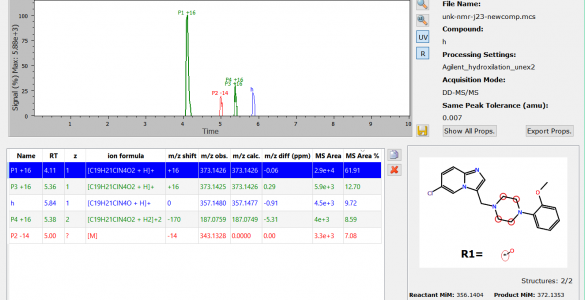
MassChemSite
MassChemSite is our new software to automatize the structural elucidation of chemical structures for high through put chemistry, late stage functionalization and in general any identification that use spectral data to do the structural elucidation in an automatic mode.
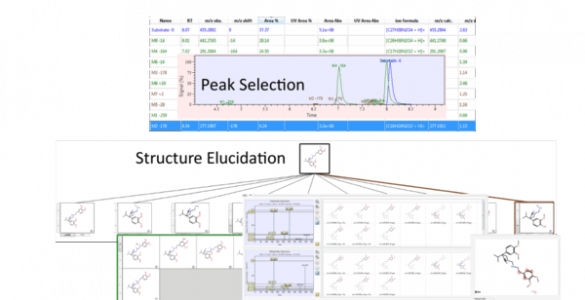
MassMetaSite
Mass-MetaSite is a new approach for the automatic identification of metabolites from Liquid Chromatography – Mass Spectrometry data, reducing manual analysis from several hours to only a few minutes per compound .
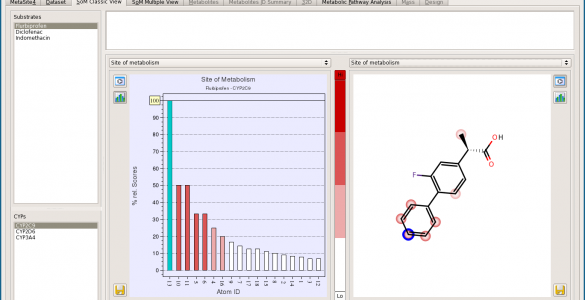
MetaSite
MetaSite is a computational procedure that predicts metabolic transformations related to cytochrome and flavin-containing monooxygenase mediated reactions in phase I metabolism. The MetaSite algorithm is unique in being the only software that is not training set dependent, and therefore exhibits improved predictive performance for novel pharmaceutical compounds.
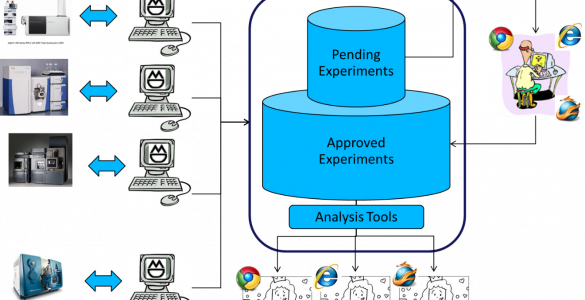
WebMetaBase
WebMetabase is a server-based application that is used for metabolite identification data storage, reviewing metabolite identification experiments, and extracting the maximum knowledge from the information loaded into the system.
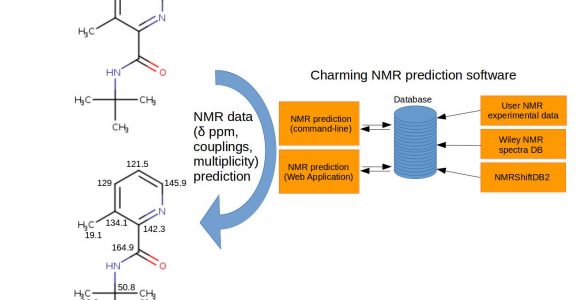
Charming NMR prediction
Charming NMR is a software engine to analyze and predict NMR properties of atoms, like chemical shift, multiplicity and coupling to other atoms. It requires a database of experimental assigned chemical shift values, which can be loaded either from NMRShiftDB2, from Wiley’s experimental NMR database or from the own user’s NMR data.
MS File Converter
This utility converts the binary files from Waters, Bruker and Thermo Fisher Scientific LCMS systems into a SQLite file format that can be used by Mass-Metasite (Molecular Discovery, Ltd).



